Grouping starred hotels in Indonesia with K-means Cluster Analysis (applied in R)
Clustering is a broad set of techniques for finding subgroups of observations within a data set. When we cluster observations, we want observations in the same group to be similar and observations in different groups to be dissimilar. Because there isn’t a response variable, this is an unsupervised method, which implies that it seeks to find relationships between the nn observations without being trained by a response variable. Clustering allows us to identify which observations are alike, and potentially categorize them therein. K-means clustering is the simplest and the most commonly used clustering method for splitting a dataset into a set of k groups.
This tutorial serves as an introduction to the k-means clustering method.
- Replication Requirements: What you’ll need to reproduce the analysis in this tutorial
- Data Preparation: Preparing our data for cluster analysis
- Clustering Distance Measures: Understanding how to measure differences in observations
- K-Means Clustering: Calculations and methods for creating K subgroups of the data
- Determining Optimal Clusters: Identifying the right number of clusters to group your data
Replication Requirements
To replicate this tutorial’s analysis you will need to load the following packages:
library(tidyverse) # data manipulation
library(cluster) # clustering algorithms
library(factoextra) # clustering algorithms & visualizationData Preparation
To perform a cluster analysis in R, generally, the data should be prepared as follows:
- Rows are observations (individuals) and columns are variables
- Any missing value in the data must be removed or estimated.
- The data must be standardized (i.e., scaled) to make variables comparable. Recall that, standardization consists of transforming the variables such that they have mean zero and standard deviation one
Here, we will use a starred hotel dataset in Indonesia consisting of one star hotel to five star hotels in 34 provinces in Indonesia
Hotel = read.delim("clipboard")
View(Hotel)
now makes the provincial variable became a column name so that it is not included for clusterization.
hotel = Hotel[,c(2:6)]
rownames(hotel) <- Hotel$Provinsi[1:34]
Cluster method itself is sensitive with the missing value in the data, so it is necessary to check the missing value first. If there is a missing value, it is necessary to handle the data, often people handle the missing value by removing these observations from the data, but this is not the best way that can be taken to handle it which is not explained in this paper. so how to know the missing value in the data ?
summary(is.na(hotel))
Based on the output there is no missing value in the data.
If the variable variables in the data have different units, then it is necessary to standardize the data first.
df <- scale(hotel[-1])
head(df)however, because the data in this test are standardized (have the same unit), there is no need for standardization of data.
Clustering Distance Measures
The classification of observations into groups requires some methods for computing the distance or the (dis)similarity between each pair of observations. The result of this computation is known as a dissimilarity or distance matrix. There are many methods to calculate this distance information; the choice of distance measures is a critical step in clustering. It defines how the similarity of two elements (x, y) is calculated and it will influence the shape of the clusters.
The choice of distance measures is a critical step in clustering. It defines how the similarity of two elements (x, y) is calculated and it will influence the shape of the clusters. The classical methods for distance measures are Euclidean and Manhattan distances, which are defined as follow:
Euclidean distance:

Manhattan distance:

Where, x and y are two vectors of length n.
Other dissimilarity measures exist such as correlation-based distances, which is widely used for gene expression data analyses. Correlation-based distance is defined by subtracting the correlation coefficient from 1. Different types of correlation methods can be used such as:
Pearson correlation distance:

Spearman correlation distance:
The spearman correlation method computes the correlation between the rank of x and the rank of y variables.


Within R it is simple to compute and visualize the distance matrix using the functions get_dist and fviz_dist from the factoextra R package. This starts to illustrate which states have large dissimilarities (red) versus those that appear to be fairly similar (teal).
get_dist: for computing a distance matrix between the rows of a data matrix. The default distance computed is the Euclidean; however,get_distalso supports distanced described in equations 2-5 above plus others.fviz_dist: for visualizing a distance matrix
distance <- get_dist(hotel[-1])
fviz_dist(distance, gradient = list(low = "#00AFBB", mid = "white", high = "#FC4E07"))
K-Means Clustering
K-means clustering is the most commonly used unsupervised machine learning algorithm for partitioning a given data set into a set of k groups (i.e. k clusters), where k represents the number of groups pre-specified by the analyst. It classifies objects in multiple groups (i.e., clusters), such that objects within the same cluster are as similar as possible (i.e., high intra-class similarity), whereas objects from different clusters are as dissimilar as possible (i.e., low inter-class similarity). In k-means clustering, each cluster is represented by its center (i.e, centroid) which corresponds to the mean of points assigned to the cluster.
The Basic Idea
The basic idea behind k-means clustering consists of defining clusters so that the total intra-cluster variation (known as total within-cluster variation) is minimized. There are several k-means algorithms available. The standard algorithm is the Hartigan-Wong algorithm (1979), which defines the total within-cluster variation as the sum of squared distances Euclidean distances between items and the corresponding centroid:

where:
- xi is a data point belonging to the cluster Ck
- μk is the mean value of the points assigned to the cluster Ck
Each observation (xi) is assigned to a given cluster such that the sum of squares (SS) distance of the observation to their assigned cluster centers (μk) is minimized.
We define the total within-cluster variation as follows:

The total within-cluster sum of square measures the compactness (i.e goodness) of the clustering and we want it to be as small as possible.
K-means Algorithm
The first step when using k-means clustering is to indicate the number of clusters (k) that will be generated in the final solution. The algorithm starts by randomly selecting k objects from the data set to serve as the initial centers for the clusters. The selected objects are also known as cluster means or centroids. Next, each of the remaining objects is assigned to it’s closest centroid, where closest is defined using the Euclidean distance (Eq. 1) between the object and the cluster mean. This step is called “cluster assignment step”. After the assignment step, the algorithm computes the new mean value of each cluster. The term cluster “centroid update” is used to design this step. Now that the centers have been recalculated, every observation is checked again to see if it might be closer to a different cluster. All the objects are reassigned again using the updated cluster means. The cluster assignment and centroid update steps are iteratively repeated until the cluster assignments stop changing (i.e until convergence is achieved). That is, the clusters formed in the current iteration are the same as those obtained in the previous iteration.
K-means algorithm can be summarized as follows:
- Specify the number of clusters (K) to be created (by the analyst)
- Select randomly k objects from the data set as the initial cluster centers or means
- Assigns each observation to their closest centroid, based on the Euclidean distance between the object and the centroid
- For each of the k clusters update the cluster centroid by calculating the new mean values of all the data points in the cluster. The centroid of a Kth cluster is a vector of length p containing the means of all variables for the observations in the kth cluster; p is the number of variables.
- Iteratively minimize the total within sum of square (Eq. 7). That is, iterate steps 3 and 4 until the cluster assignments stop changing or the maximum number of iterations is reached. By default, the R software uses 10 as the default value for the maximum number of iterations.
Computing k-means clustering in R
We can compute k-means in R with the kmeans function. Here will group the data into two clusters (centers = 2). The kmeans function also has an nstart option that attempts multiple initial configurations and reports on the best one. For example, adding nstart = 25 will generate 25 initial configurations. This approach is often recommended.
k2 <- kmeans(hotel, centers = 2, nstart = 25)
str(k2)
## List of 9
## $ cluster : Named int [1:34] 1 2 2 2 1 2 1 1 ...
## ..- attr(*, "names")= chr [1:534 "1" "2" "3" "4" ...
## $ centers : num [1:2, 1:4] 2.36 55.08 43.78 ...
## ..- attr(*, "dimnames")=List of 2
## .. ..$ : chr [1:2] "1" "2"
## .. ..$ : chr [1:5] "Start.5" "Start.4" "Start.3" "Start.2"
## $ totss : num 4981
## $ withinss : num [1:2] 17944 6575
## $ tot.withinss: num 24519
## $ betweenss : num 25322
## $ size : int [1:2] 15 19
## $ iter : int 1
## $ ifault : int 0
## - attr(*, "class")= chr "kmeans"cluster: A vector of integers (from 1:k) indicating the cluster to which each point is allocated.centers: A matrix of cluster centers.totss: The total sum of squares.withinss: Vector of within-cluster sum of squares, one component per cluster.tot.withinss: Total within-cluster sum of squares, i.e. sum(withinss).betweenss: The between-cluster sum of squares, i.e. $totss-tot.withinss$.size: The number of points in each cluster.
If we print the results we’ll see that our groupings resulted in 2 cluster sizes of 15 and 19. We see the cluster centers (means) for the two groups across the four variables (1 = Aceh, 2 = Sumatra Utara, 3 = Sumatra Barat, 4 = Riau). We also get the cluster assignment for each observation (i.e. Aceh was assigned to cluster 1, Sumatra Utara was assigned to cluster 2, etc.).
k2
We can also view our results by using fviz_cluster. This provides a nice illustration of the clusters. If there are more than two dimensions (variables) fviz_cluster will perform principal component analysis (PCA) and plot the data points according to the first two principal components that explain the majority of the variance.
fviz_cluster(k2, data = hotel)
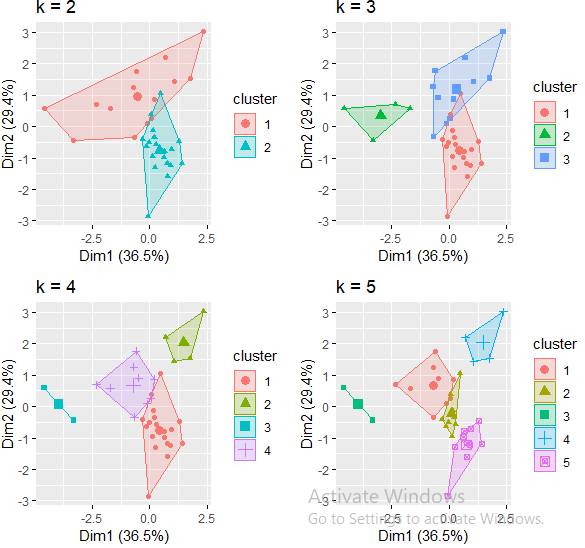
Because the number of clusters (k) must be set before we start the algorithm, it is often advantageous to use several different values of k and examine the differences in the results. We can execute the same process for 3, 4, and 5 clusters, and the results are shown in the figure:
k3 <- kmeans(hotel, centers = 3, nstart = 25)
k4 <- kmeans(hotel, centers = 4, nstart = 25)
k5 <- kmeans(hotel, centers = 5, nstart = 25)# plots to compare
p1 <- fviz_cluster(k2, geom = "point", data = hotel[-1]) + ggtitle("k = 2")
p2 <- fviz_cluster(k3, geom = "point", data = hotel[-1]) + ggtitle("k = 3")
p3 <- fviz_cluster(k4, geom = "point", data = hotel[-1]) + ggtitle("k = 4")
p4 <- fviz_cluster(k5, geom = "point", data = hotel[-1]) + ggtitle("k = 5")library(gridExtra)
grid.arrange(p1, p2, p3, p4, nrow = 2)

Although this visual assessment tells us where true dilineations occur (or do not occur such as clusters 2 & 4 in the k = 5 graph) between clusters, it does not tell us what the optimal number of clusters is.
Determining Optimal Clusters
As you may recall the analyst specifies the number of clusters to use; preferably the analyst would like to use the optimal number of clusters. To aid the analyst, the following explains the three most popular methods for determining the optimal clusters, which includes:
Elbow Method
Recall that, the basic idea behind cluster partitioning methods, such as k-means clustering, is to define clusters such that the total intra-cluster variation (known as total within-cluster variation or total within-cluster sum of square) is minimized:

where Ck is the kthkth cluster and W(Ck) is the within-cluster variation. The total within-cluster sum of square (wss) measures the compactness of the clustering and we want it to be as small as possible. Thus, we can use the following algorithm to define the optimal clusters:
- Compute clustering algorithm (e.g., k-means clustering) for different values of k. For instance, by varying k from 1 to 10 clusters
- For each k, calculate the total within-cluster sum of square (wss)
- Plot the curve of wss according to the number of clusters k.
- The location of a bend (knee) in the plot is generally considered as an indicator of the appropriate number of clusters.
We can implement this in R with the following code. The results suggest that 3 is the optimal number of clusters as it appears to be the bend in the knee (or elbow).
Fortunately, this process to compute the “Elbow method” has been wrapped up in a single function (fviz_nbclust):
set.seed(123)
fviz_nbclust(hotel, kmeans, method = "wss")
Average Silhouette Method
In short, the average silhouette approach measures the quality of a clustering. That is, it determines how well each object lies within its cluster. A high average silhouette width indicates a good clustering. The average silhouette method computes the average silhouette of observations for different values of k. The optimal number of clusters k is the one that maximizes the average silhouette over a range of possible values for k
We can use the silhouette function in the cluster package to compuate the average silhouette width. The following code computes this approach for 1-15 clusters. The results show that 2 clusters maximize the average silhouette values with 4 clusters coming in as second optimal number of clusters.
fviz_nbclust(hotel, kmeans, method = "silhouette")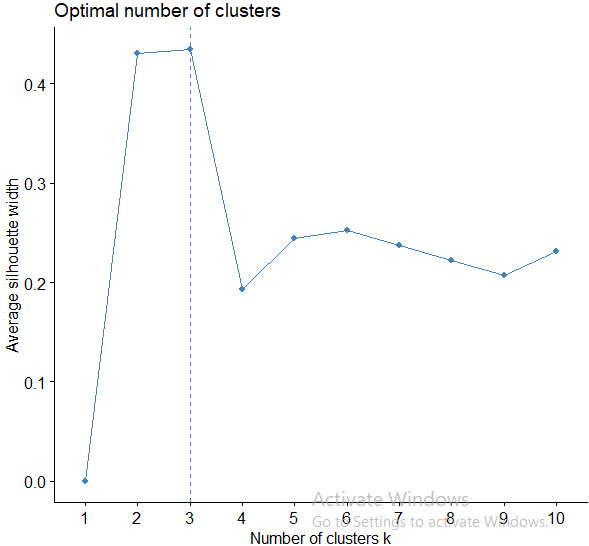
Gap Statistic Method
The gap statistic has been published by R. Tibshirani, G. Walther, and T. Hastie (Standford University, 2001). The approach can be applied to any clustering method (i.e. K-means clustering, hierarchical clustering). The gap statistic compares the total intracluster variation for different values of k with their expected values under null reference distribution of the data (i.e. a distribution with no obvious clustering). The reference dataset is generated using Monte Carlo simulations of the sampling process. That is, for each variable (xi) in the data set we compute its range [min(xi),max(xj)][min(xi),max(xj)] and generate values for the n points uniformly from the interval min to max.
To compute the gap statistic method we can use the clusGap function which provides the gap statistic and standard error for an output.
# compute gap statistic
set.seed(123)
gap_stat <- clusGap(df, FUN = kmeans, nstart = 25,
K.max = 10, B = 50)
# Print the result
print(gap_stat, method = "firstmax")# Visual
fviz_gap_stat(gap_stat)

Extracting Results
With most of these approaches suggesting 3 as the number of optimal clusters, we can perform the final analysis and extract the results using 3 clusters.
set.seed(123)
final <- kmeans(hotel, 3, nstart = 25)
print(final)We can visualize the results using fviz_cluster:
fviz_cluster(final, data = hotel[-1])
Conclussion
Next, we will display the data frame of the test data that has been added with the cluster results as follows
df.cluster = data.frame(hotel, final$cluster)
View(df.cluster)

we will see the characteristics of each existing cluster as follows :
k1.1 = subset(df.cluster, k2.cluster == 1)
k1.2 = subset(df.cluster, k2.cluster == 2)
k1.3 = subset(df.cluster, k2.cluster == 3)meank1.1 = sapply(k1.1, mean)
meank1.2 = sapply(k1.2, mean)
meank1.3 = sapply(k1.3, mean)mean.tot = rbind(meank1.1, meank1.2, meank1.3)
barplot(t(mean.tot[,-6]), beside = TRUE)


Based on the graph we know that :
- In cluster one consists of provinces that have the best quality both of 4-star hotels and 3-star hotels compared to cluster two, but do not have 5-star hotels and have the lowest 1-star hotel services compared to cluster 2 and cluster 3.
- In cluster two consists of provinces that have the best quality of 1-star hotels and 2-star hotels compared to clusters one and three, but have the lowest quality of 3-star hotels and 4-star hotels but have the best 5-star hotels compared to cluster one
- In cluster three consists of provinces that have the best quality both of 5-star hotels and 3-star hotels compared to clusters one and cluster two, and have quality 2-star hotels that are almost as good as cluster 2, and have the best 1-star hotel quality compared to cluster 1.
Additional Comments
K-means clustering is a very simple and fast algorithm. Furthermore, it can efficiently deal with very large data sets. However, there are some weaknesses of the k-means approach.
One potential disadvantage of K-means clustering is that it requires us to pre-specify the number of clusters. Hierarchical clustering is an alternative approach which does not require that we commit to a particular choice of clusters. Hierarchical clustering has an added advantage over K-means clustering in that it results in an attractive tree-based representation of the observations, called a dendrogram. A future tutorial will illustrate the hierarchical clustering approach.
An additional disadvantage of K-means is that it’s sensitive to outliers and different results can occur if you change the ordering of your data. The Partitioning Around Medoids (PAM) clustering approach is less sensititive to outliers and provides a robust alternative to k-means to deal with these situations. A future tutorial will illustrate the PAM clustering approach.
References :
